Solve Global Optimization Problems in MATLAB with differential_evolution()
Part 8 of the Python Is The Ultimate MATLAB Toolbox series.
- Introduction
- Differential evolution
- An optimization problem: shape alignment
- Setting up the cost function
- Visualizing the cost function
- Finding the global minimum
- Python solution with
scipy.optimize.differential_evolution() - Additional Python optimization packages
Introduction
MATLAB’s fminsearch() function finds local minima of unbounded
multivariable functions.
With a lucky starting guess, it may even return
the global minimum.
For more rigirous searches of global minima, or control over
the search domain, you’ll need more powerful options such as those
in the
Optimization Toolbox.
Alternatives to the Optimization Toolbox include global optimizers from
- the FileExchange, specifically a differential evolution solver as used in this post, or
- the
scipy.optimizemodule, provided that you’re willing to include Python in your MATLAB workflow and implement your objective function in Python.
The first option is the most convenient for MATLAB users as it provides a pure MATLAB solution. The second option may be necessary in corporate networks that restrict access to the Internet but include Python on the same computers that have MATLAB. This post covers both options.
Differential evolution
scipy.optimize has three global optimizers: shgo(),
dual_annealing(), and differential_evolution().
Of the three, the most
powerful is arguably differential_evolution() which implements
the Storn and Price algorithm1 of the same name.
Briefly, the differential evolution algorithm works by starting with a population of random guesses to the solution. Guesses which yield better answers to the cost function are combined to form new guesses while poorly performing guesses are eliminated. The algorithm repeats this process until the guesses converge.
An optimization problem: shape alignment
Say you have two images of a triangular widget, where the widget’s position in one image is offset from the position in the other. After some image processing, perhaps with OpenCV to identify only the corners, you’re left with these two images:
Target image on the left, known image on the right.
Our goal is to figure out how far to translate the shape on the right until it overlaps the one on the left (we’ll ignore rotation and assume the widgets are always oriented the same way). Simple, right? Just compute the horizontal and vertical differences of a common corner such as the upper tip:
However if the widget on the left is on a noisy background and the corner identifcation code returns many spurious points such as shown here,
then problem is much harder since there’s no clear way to determine which known point corresponds to which target point.
There exist algorithms that can compute accurate offsets quickly but here we’ll take a lazy approach and let an optimizer figure this out. The cost function can be the sum of distances from each corner of our known image to the nearest point on the target image. This sum should be zero if all three corners of the known image perfectly overlay the three nearest target points.
We’ll start with three known points on a unit grid. To this we’ll add ten target points which are made of three copies of the known points plus an offset of (-0.4, -0.2) and seven additional random points. We want an optimizer to return the offset using only the locations of the original three known points and the ten target points. The code below makes a 10 x 2 array of random coordinates for the targets then arbitrarily replaces the 3rd, 6th, and 10th points with the known points plus the offset.
Python:
In : import numpy as np
In : known = np.array([[0.6, 0.5], [0.6, 0.9], [0.7,0.5]])
In : target = np.random.rand(10,2)
In : truth_offset = np.array([-0.4, -0.2])
In : target[2,:] = known[0,:] + truth_offset
In : target[5,:] = known[1,:] + truth_offset
In : target[9,:] = known[2,:] + truth_offsetMATLAB:
>> known = [0.6 0.5; 0.6 0.9; 0.7 0.5];
>> target = rand(10,2);
>> truth_offset = [-0.4 -0.2];
>> target( 3,:) = known(1,:) + truth_offset;
>> target( 6,:) = known(2,:) + truth_offset;
>> target(10,:) = known(3,:) + truth_offset;Python and MATLAB will come up with different random target points which will make it harder to compare solutions. To keep things consistent I’ll run the MATLAB version to produce a set of values, then hard-code these into both MATLAB and Python test environments:
Python:
known = np.array([[0.6, 0.5], [0.6, 0.9], [0.7,0.5]])
target = np.array([[ 8.147236863931789e-01, 1.576130816775483e-01],
[ 9.057919370756192e-01, 9.705927817606157e-01],
[ 2.0e-01 , 3.0e-01 ],
[ 9.133758561390194e-01, 4.853756487228412e-01],
[ 6.323592462254095e-01, 8.002804688888001e-01],
[ 2.0e-01 , 7.0e-01 ],
[ 2.784982188670484e-01, 4.217612826262750e-01],
[ 5.468815192049838e-01, 9.157355251890671e-01],
[ 9.575068354342976e-01, 7.922073295595544e-01],
[ 3.0e-01 , 3.0e-01 ]])MATLAB:
>> known = [0.6 0.5; 0.6 0.9; 0.7 0.5];
>> target = [ 8.147236863931789e-01, 1.576130816775483e-01;
9.057919370756192e-01, 9.705927817606157e-01;
2.0e-01 , 3.0e-01 ;
9.133758561390194e-01, 4.853756487228412e-01;
6.323592462254095e-01, 8.002804688888001e-01;
2.0e-01 , 7.0e-01 ;
2.784982188670484e-01, 4.217612826262750e-01;
5.468815192049838e-01, 9.157355251890671e-01;
9.575068354342976e-01, 7.922073295595544e-01;
3.0e-01 , 3.0e-01 ];Next we need to find the closest target point for each known point To do that we’ll need to compute distances from all targets to all knowns then pick the target with the smallest distance for each known point. The pairwise distance matrix computation is simple with NumPy arrays in Python and matrices in MATLAB because both support broadcasting. With broadcasting, subtracting a row vector with nR terms from a column vector with nC terms leaves us with an nR x nC matrix of all pairwise differences:
MATLAB:
>> format short e
>> known(:,1)'
6.0000e-01 6.0000e-01 7.0000e-01
>> target(:,1)
8.1472e-01
9.0579e-01
2.0000e-01
9.1338e-01
6.3236e-01
2.0000e-01
2.7850e-01
5.4688e-01
9.5751e-01
3.0000e-01then
MATLAB:
>> target(:,1) - known(:,1)'
2.1472e-01 2.1472e-01 1.1472e-01
3.0579e-01 3.0579e-01 2.0579e-01
-4.0000e-01 -4.0000e-01 -5.0000e-01
3.1338e-01 3.1338e-01 2.1338e-01
3.2359e-02 3.2359e-02 -6.7641e-02
-4.0000e-01 -4.0000e-01 -5.0000e-01
-3.2150e-01 -3.2150e-01 -4.2150e-01
-5.3118e-02 -5.3118e-02 -1.5312e-01
3.5751e-01 3.5751e-01 2.5751e-01
-3.0000e-01 -3.0000e-01 -4.0000e-01The same can be done with NumPy arrays but there one must explicitly
add a second dimension to one of the two column extracts
with np.newaxis.
Without this step, the arrays of x coordinates are merely
one dimensional arrays with no concept of row or column orientation.
(A detailed explanation can be found in section 11.1.12 of my
book.)
Python:
In : target[:,0][:,np.newaxis] - known[:,0]
Out:
array([[ 0.21472369, 0.21472369, 0.11472369],
[ 0.30579194, 0.30579194, 0.20579194],
[-0.4 , -0.4 , -0.5 ],
[ 0.31337586, 0.31337586, 0.21337586],
[ 0.03235925, 0.03235925, -0.06764075],
[-0.4 , -0.4 , -0.5 ],
[-0.32150178, -0.32150178, -0.42150178],
[-0.05311848, -0.05311848, -0.15311848],
[ 0.35750684, 0.35750684, 0.25750684],
[-0.3 , -0.3 , -0.4 ]])By extending the broadcasting idea to include the delta y computation then squaring and summing, we can get the complete 10 x 3 array of squares of distances between points with just one (long) line of code:
Python:
In : dist2 = ( target[:,0][:,np.newaxis] - known[:,0] )**2 + \
: ( target[:,1][:,np.newaxis] - known[:,1] )**2
In : dist2
Out:
array([[0.16333506, 0.5972446 , 0.13039033],
[0.31496628, 0.09849205, 0.26380789],
[0.2 , 0.52 , 0.29 ],
[0.0984183 , 0.27011778, 0.04574313],
[0.09121548, 0.01099111, 0.09474363],
[0.2 , 0.2 , 0.29 ],
[0.10948469, 0.33207567, 0.18378505],
[0.1756576 , 0.00306918, 0.1962813 ],
[0.21319626, 0.1394304 , 0.15169489],
[0.13 , 0.45 , 0.2 ]])MATLAB:
>> dist2 = ( target(:,1) - known(:,1)' ).^2 + ...
( target(:,2) - known(:,2)' ).^2
1.6334e-01 5.9724e-01 1.3039e-01
3.1497e-01 9.8492e-02 2.6381e-01
2.0000e-01 5.2000e-01 2.9000e-01
9.8418e-02 2.7012e-01 4.5743e-02
9.1215e-02 1.0991e-02 9.4744e-02
2.0000e-01 2.0000e-01 2.9000e-01
1.0948e-01 3.3208e-01 1.8379e-01
1.7566e-01 3.0692e-03 1.9628e-01
2.1320e-01 1.3943e-01 1.5169e-01
1.3000e-01 4.5000e-01 2.0000e-01meaning, for example, the square of the distance between the 8th target point (== 8th row) and 3rd known point (== 3rd column) is 1.9628e-01. All that remains is to take the square root of every term.
Setting up the cost function
Now that we know how far apart each target point is from each known point, we just need to find the closest ones then sum their distances. Taking the minumum over each column, then summing gives us the cost function:
Python:
In : dist = np.sqrt( ( target[:,0][:,np.newaxis] - known[:,0] )**2 +
: ( target[:,1][:,np.newaxis] - known[:,1] )**2 )
In : np.min(dist, axis=0)
Out: array([0.30201901, 0.05540018, 0.21387643])
In : np.min(dist, axis=0).sum()
Out: 0.5712956164197415MATLAB:
>> dist = sqrt( ( target(:,1) - known(:,1)' ).^2 + ...
( target(:,2) - known(:,2)' ).^2 );
>> min(dist)
0.3020 0.0554 0.2139
>> sum(min(dist))
0.5713Optimizers need to call the function that’s to be minimized
so we’ll rewrite the expressions above as functions called cost()2.
We merely need to wrap the distance summation computation
into a function:
Python:
import numpy as np
def cost(known, target):
dist = np.sqrt( ( target[:,0][:,np.newaxis] - known[:,0] )**2 +
( target[:,1][:,np.newaxis] - known[:,1] )**2 )
return np.min(dist, axis=0).sum()MATLAB:
function [sum_dist] = cost(known, target)
dist = sqrt( ( target(:,1) - known(:,1)' ).^2 + ...
( target(:,2) - known(:,2)' ).^2 );
sum_dist = sum(min(dist));
endVisualizing the cost function
It’s instructive to see how the cost function varies across the search domain but this is only practical if the cost function has two or fewer independent variables and is inexpensive to compute. The cost function chosen for this post was obviously tailored to meet these criteria. I’d have liked to allow the target shape to be oriented at an arbitrary angle but then the offset would consist of three independent variables, x, y, and phi, and the cost function would need to be rendered as a 3D cube. That’s possible but requires an advanced visualization tool that supports cutting planes.
Here are functions to display a contour plot of the cost function across the search domain. Both functions descretize the search domain of -1 <= x <= 1 and -1 <= y <= 1 into an evenly spaced grid of 100 x 100 points then compute the cost function at each of these. Afterwards we’ll view the 100 x 100 cost terms on a contour plot.
Python:
import numpy as np
import matplotlib.pyplot as plt
def plot_cost(known, target): # {{{
N = 100
[offset_x, offset_y] = np.meshgrid(np.linspace(-1,1,N),np.linspace(-1,1,N))
C = np.zeros_like(offset_x) # cost at each offset point
offset_target = np.zeros_like(target)
for r in range(N):
for c in range(N):
offset = np.array([offset_x[r,c], offset_y[r,c]])
offset_target = target + offset
C[r,c] = cost(known, offset_target)
fig, ax = plt.subplots()
plt.contourf(offset_x, offset_y, C, levels=20)
ax.set_aspect('equal')
ax.set_xlim([-1,1.0])
ax.set_xlabel('X offset')
ax.set_ylabel('Y offset')
ax.set_ylim([-1,1.0])
plt.colorbar()
plt.show()MATLAB:
function plot_cost(known, target)
N = 100;
[offset_x, offset_y] = meshgrid(linspace(-1,1,N), linspace(-1,1,N));
C = zeros(size(offset_x)); % cost at each offset point
offset_target = zeros(size(target));
for r = 1:N
for c = 1:N
offset = [offset_x(r,c), offset_y(r,c)];
offset_target = target + offset;
C(r,c) = cost(known, offset_target);
end
end
levels = 20;
contourf(offset_x, offset_y, C, levels)
colorbar()
endFor our first plot let’s start with the easy case with just three target points which are offset from three known points:
Python:
import numpy as np
import matplotlib.pyplot as plt
known = np.array([[0.6, 0.5], [0.6, 0.9], [0.7,0.5]])
truth_offset = np.array([-0.4, -0.2])
target = known + truth_offset;
plot_cost(known, target)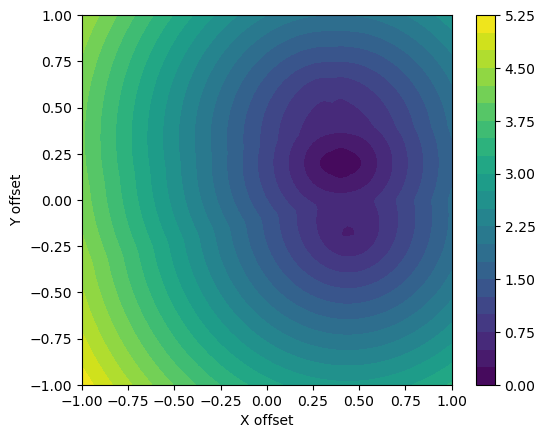
Cost function for three targets, Python.
There’s a fairly distinct minimum around (0.4, 0.2). This is the amount
we’d need to add to our target points to make them overlay the known
points. Looking at it from the other direction,
(-0.4, -0.2) was added to our known points to produce the targets.
The cost function is fairly smooth with a clear minimum and should
not present much challenge for gradient-based optimizers
such as MATLAB’s fminsearch().
Now let’s plot the harder case of 10 target points—3 from shifting the known points, plus the 7 noise points:
Python:
import numpy as np
import matplotlib.pyplot as plt
known = np.array([[0.6, 0.5], [0.6, 0.9], [0.7,0.5]])
target = np.array([[ 8.147236863931789e-01, 1.576130816775483e-01],
[ 9.057919370756192e-01, 9.705927817606157e-01],
[ 2.0e-01 , 3.0e-01 ],
[ 9.133758561390194e-01, 4.853756487228412e-01],
[ 6.323592462254095e-01, 8.002804688888001e-01],
[ 2.0e-01 , 7.0e-01 ],
[ 2.784982188670484e-01, 4.217612826262750e-01],
[ 5.468815192049838e-01, 9.157355251890671e-01],
[ 9.575068354342976e-01, 7.922073295595544e-01],
[ 3.0e-01 , 3.0e-01 ]])
plot_cost(known, target)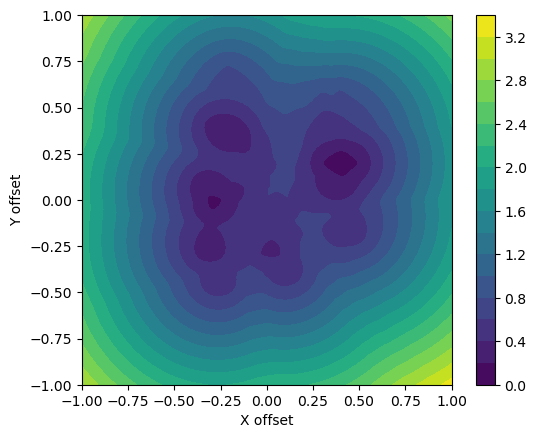
Cost function for ten targets, Python.
This time the global minimum is hard to identify. If we add even more noise points the plot will have more local minima and the optimization problem becomes even more challenging.
For completeness here are the MATLAB versions of the two plots:
MATLAB:
known = [0.6 0.5; 0.6 0.9; 0.7 0.5];
truth_offset = [-.4 -.2];
target3 = known + truth_offset;
plot_cost(known, target3)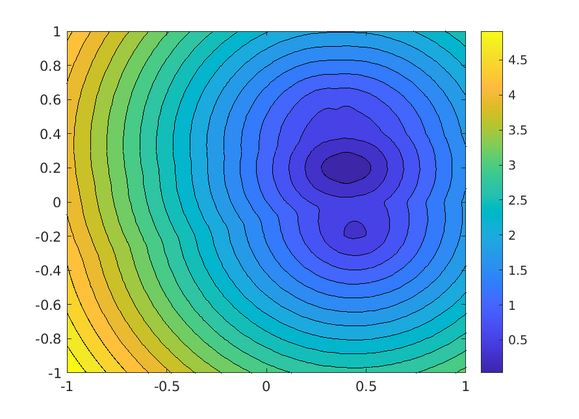
Cost function for three targets, MATLAB.
MATLAB:
known = [0.6 0.5; 0.6 0.9; 0.7 0.5];
target = [ 8.147236863931789e-01, 1.576130816775483e-01;
9.057919370756192e-01, 9.705927817606157e-01;
2.0e-01 , 3.0e-01 ;
9.133758561390194e-01, 4.853756487228412e-01;
6.323592462254095e-01, 8.002804688888001e-01;
2.0e-01 , 7.0e-01 ;
2.784982188670484e-01, 4.217612826262750e-01;
5.468815192049838e-01, 9.157355251890671e-01;
9.575068354342976e-01, 7.922073295595544e-01;
3.0e-01 , 3.0e-01 ];
plot_cost(known, target)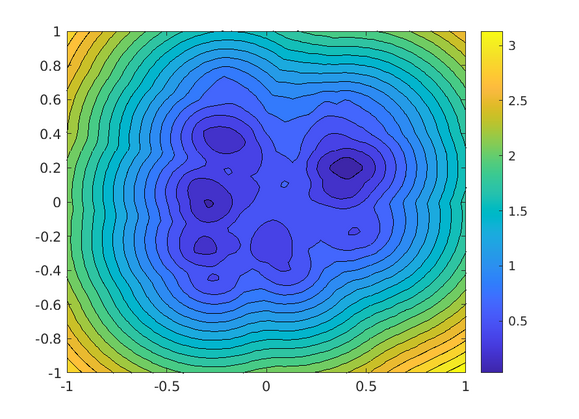
Cost function for ten targets, MATLAB.
Finding the global minimum
We’re ready to put our optimizers to work.
Before we jump to differential evolution, let’s try the fminsearch()
function in MATLAB, first with the three target points:
MATLAB:
>> start_offset = [0 0];
>> fn = @(offset) cost(known, target3 + offset);
>> fminsearch(fn, start_offset)
0.4000 0.2000Looks good. Now the much harder case with the ten target points:
MATLAB:
>> start_offset = [0 0];
>> fn = @(offset) cost(known, target + offset);
>> fminsearch(fn, start_offset)
0.0285 -0.2725Not so good—(0.0285, -0.2725) is one of the several local minima.
The Optimization Toolbox has functions to find the global minimum
but if that’s not available, a good alternative is a
differential evolution based solver from the FileExchange
or from scipy.optimize.
Python solution with scipy.optimize.differential_evolution()
SciPy’s differential_evolution() calling arguments are similar to
MATLAB’s fminsearch() but include an additional array of
bounds on the independent variables (the x and y values of
the offset). The first argument is a
handle to a function, we’ll call it fn() to match the @fn()
anonymous function used in MATLAB since it serves the same purpose—in
both Python and MATLAB, fn calls our cost function with
trial values for independent variables plus other required
arguments, namely the known and target arrays, passed
through with the keyword argument args. Here’s what that
looks like:
Python:
def fn(independent_vars, *data):
x_offset, y_offset = independent_vars
known, target = data
offset_target = target + np.array([x_offset, y_offset])
return cost(known, offset_target)
bounds=([-1.0, 1.0], [ -1.0, 1.0])
result = differential_evolution(fn, bounds, args=(known, target))Solutions for the three and ten point target cases return the correct global minimum for both cases:
Python:
In : differential_evolution(fn, bounds, args=(known, target3))
Out:
fun: 4.052626611931048e-16
message: 'Optimization terminated successfully.'
nfev: 3033
nit: 98
success: True
x: array([0.4, 0.2])
In : differential_evolution(fn, bounds, args=(known, target))
Out:
fun: 3.3664710476199897e-16
message: 'Optimization terminated successfully.'
nfev: 2913
nit: 94
success: True
x: array([0.4, 0.2])Differential evolution involves random guesses for the independent
variables so the number of function evaluations (nfev in the
solution structure) varies between runs.
Oddly, in this instance, the easier case of three target points required more
evaluations than the harder case of ten points.
If the cost function is expensive—for example a structural analysis
run using Ansys or NASTRAN to find the maximum stress in
a finite element model—you may want to modify the
differential_evolution() optional parameters
to find a suitable solution with fewer evaluations, or to
run on multiple processors.
Individual elements of the returned structure are accessed through dot notation, for example,
Python:
In : result = differential_evolution(fn, bounds, args=(known, target3))
In : result.nfev
Out: 3213
In : result.x
Out: array([0.4, 0.2])The complete Python program with all code above is optim_w_DE.py.
MATLAB solution with differential_evolution() from the FileExchange
Markus Buehren
posted a thorough implementation of
differential evolution
entirely in MATLAB on the
FileExchange.
I didn’t spend enough time with it to learn how to pass arbitrary inputs
to the cost function so the implementation below embeds the known
and target arrays directly in cost().
I expanded the zip file to the directory /home/al/matlab/DE
on my computer.
To use Markus’s solver
this directory must be in the MATLAB search path so the
script below begins with an addpath() to this location.
MATLAB: optim_w_fileexchange.m
|
|
Running this script yields the correct solution
MATLAB:
:
'Shape alignment' finished after given maximum number of 50 iterations.
Elapsed time: 0.37 seconds
Number of function evaluations: 961
Mean function evaluation time: 0.0003928 seconds
Final result after 50 iteration(s).
Best member:
parameter1 = 0.4;
parameter2 = 0.2;
Evaluation value: 5.10014e-14
Results are saved in file cost_result_01.mat.and additionally brings up plots and writes .mat files
to the working directory.
MATLAB solution with scipy.optimize.differential_evolution()
If the FileExchange implementation of differential evolution isn’t
an option for you, use the implementation in SciPy instead.
Although
scipy.optimize.differential_evolution() can’t be called from MATLAB
directly—it requires a Python evaluation function, the fn()
mentioned above—the
call can be put in a Python wrapper and the wrapper called from MATLAB.
This means, however, the cost function must also be implemented in Python
(or I suppose could use the
MATLAB engine in Python
to call the MATLAB cost function from Python although I’ve never attempted this).
Here’s what the wrapper looks like for our problem:
Python: diffev_bridge.py
|
|
The MATLAB invocation to this wrapper is
MATLAB (>= 2020b): optim_w_python.m
|
|
running it returns a solution dictionary for the three point and ten point cases:
MATLAB (>= 2020b):
>> optim_w_python
result =
Python dict with no properties.
{'success': True,
'message': 'Optimization terminated successfully.',
'x': array([0.4, 0.2]),
'nfev': 3153}
result =
Python dict with no properties.
{'success': True,
'message': 'Optimization terminated successfully.',
'x': array([0.4, 0.2]),
'nfev': 3033}Individual items from the solution dictionary can be accessed
using the dictionary .get() method. Alternatively, convert
the dictionary to a native MATLAB struct with
py2mat.m:
MATLAB (>= 2020b):
>> offset = double(result.get('x'))
0.4000 0.2000
>> soln = py2mat(result)
struct with fields:
success: 1
message: "Optimization terminated successfully."
x: [0.4000 0.2000]
nfev: 2943
>> soln.x
0.4000 0.2000Avoiding MATLAB crashes on Linux
MATLAB 2020b on Ubuntu 20.04 may crash
at diffev.call_DE() when running optim_w_python.m.
The solution is to start MATLAB from a
suitably-configured Python environment
and to include these lines in startup.m:
MATLAB (>= 2020b):
py.sys.setdlopenflags(int32(10));
os = py.importlib.import_module('os');
os.environ.update(pyargs('KMP_DUPLICATE_LIB_OK','TRUE'))Additional Python optimization packages
Optimization is a massive topic that spans problem classes that differential evolution cannot solve. See Keivan Tafakkori’s list of optimization packages for Python for an overview of additional solvers.
Table of Contents | Previous: Interact with Redis | Next : Solve simultaneous nonlinear equations
-
Storn, R.; Price, K. (1997). “Differential evolution - a simple and efficient heuristic for global optimization over continuous spaces”. Journal of Global Optimization. 11 (4): 341–359. doi:10.1023/A:1008202821328. S2CID 5297867. ↩︎
-
If you are using the Audio Toolbox, Communications Toolbox, or DSP System Toolbox you’ll want to call the function something other than
cost()since these toolboxes also have acost()function. ↩︎
